Abstract
Introduction:
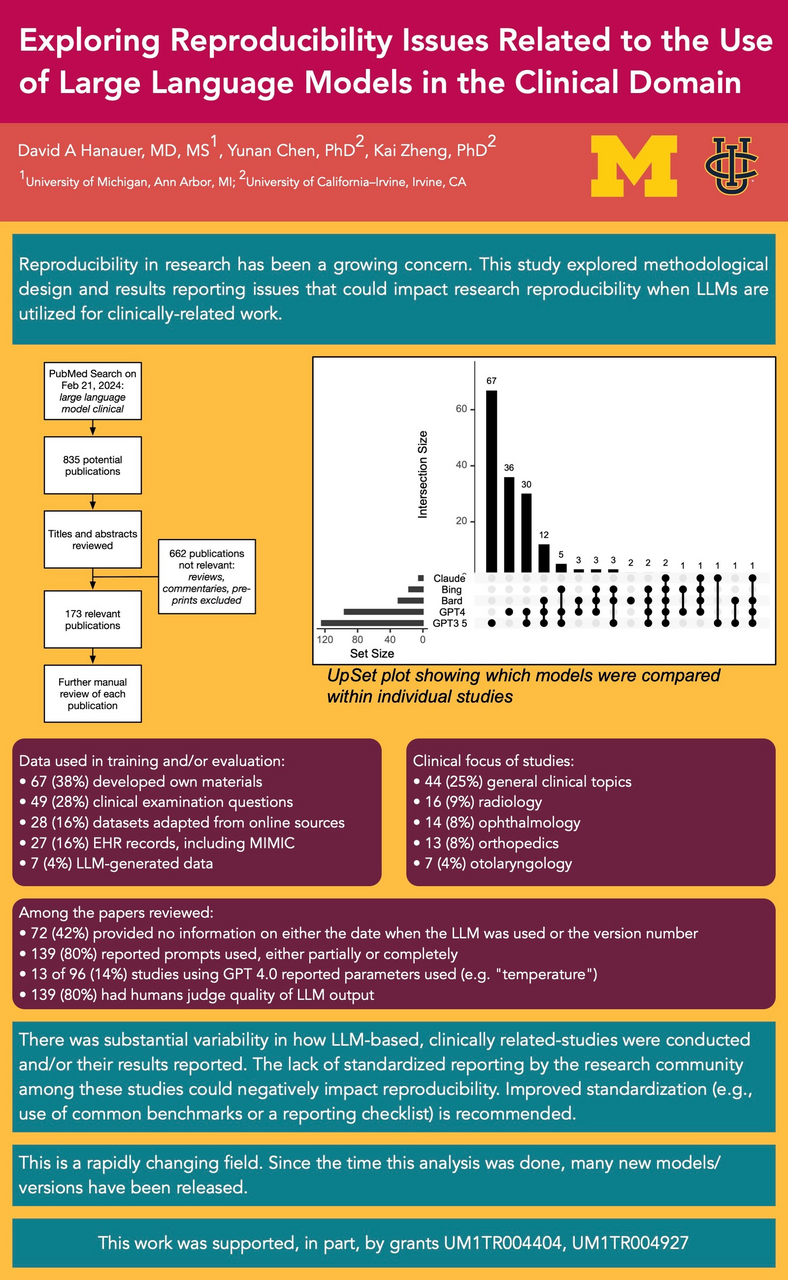
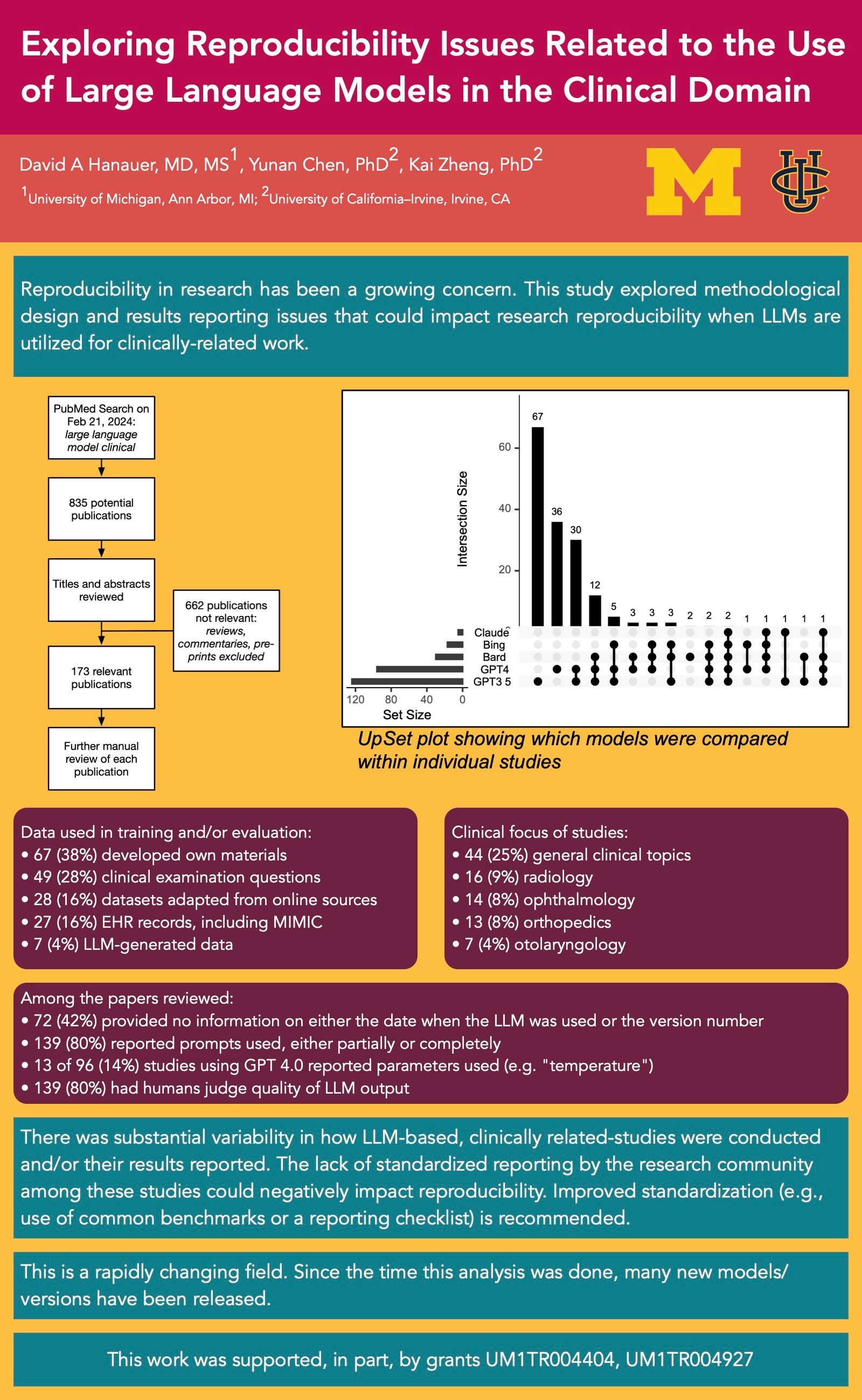
Reproducibility in research, or the lack thereof, has been a growing concern in the scientific community, contributed by factors such as “selective reporting” and “methods, code unavailable”, among others. 1 The recent advances in, and easy access to, large language models (LLMs) have led to numerous studies that applied LLMs to clinically related topics. The goal of this work was to identify and quantify methodological design and results reporting issues that could impact research reproducibility when LLMs are involved.
Methods:
We searched PubMed on February 21, 2024, using the query “large language model clinical,” with the search scope limited to papers published in English and within the prior year. Specific names of LLMs were not used in the query to reduce bias. Studies using non-publicly available LLMs were excluded. Also excluded were reviews, commentaries, and non-peer-reviewed manuscripts from pre-print archives. Metrics were chosen based on study factors researchers could easily access or report (e.g., model name, version number) and that would be important for study reproducibility.
Results:
The literature search yielded a total of 835 publications. After screening their titles and abstracts, 173 were deemed meeting the criteria and were included in the review. ChatGPT 3.5 was the most commonly studied LLM (n=124; 72%), followed by ChatGPT 4.0 (n=96; 55%), Bard (n=30; 17%), Bing (n=17; 10%), and Claude (n=5; 3%). Of note, 31 papers (18%) only reported using “ChatGPT” without specifying the version. In such cases, we classified them as using ChatGPT 3.5, which was the primary freely accessible LLM in use at the time. Many studies evaluated multiple LLMs to comparatively assess their performance, as shown in the Figure. With respect to empirical data used in LLM training and/or evaluation, 67 studies (39%) developed their own materials, followed by using clinical examination questions (n=49; 28%), datasets adapted from online sources (n=28; 16%), and electronic health records data including MIMIC (n= 27; 16%). In 7 studies (4%), the researchers generated new materials from the LLM without using any input data. A substantial proportion of the studies (n=72, 42%) provided no information on either the date when the LLM was used or the version number. Prompts used were reported either partially or completely in 139 papers (80%). Of the 96 papers using GPT 4.0, which supports fine-tuning through user-specified settings (e.g., “temperature”), only 13 studies (14%) reported the parameters used. Human annotators were involved in 139 studies to judge the quality of LLM output. Of these studies, 99 (71%) reported involving ≥ 2 annotators; 30 (22%) did not specify. Further results continued on the poster.
Discussion and Conclusions:
Our findings show that there was substantial variability in how LLM-based, clinically related-studies were conducted and/or their results reported. For example, many papers failed to disclose the version of the LLM involved and the parameters used, which could have significant implications on reproducibility. Variability in the training (i.e., clinical expertise) of annotators in studies could also have an impact when interpreting clinical relevance/accuracy of the LLM. The lack of standardized reporting by the research community among these studies could negatively impact reproducibility. Improved standardization (e.g., use of common benchmarks or a reporting checklist) is recommended.